 Such blocks are obtained after removing unaligned regions, and then splitting contigs at misassembly breakpoints. It is calculated using aligned blocks instead of contigs. NGA50, a reference-aware version of N50 metric.Number of genes in the assembly, completely or partially covered, based on a user-provided list of gene positions in the reference.This occurs due to multiple reasons, including overestimating repeat multiplicities and overlaps between contigs. If the assembly contains many contigs that cover the same regions, its duplication ratio will significantly exceed 1. Duplication ratio, the total number of aligned bases in the assembly divided by the total number of those in the reference.Genome fraction %, assembled part of the reference.Numbers of mismatches and indels, over the assembly and per 100 kb.Number and total length of unaligned contigs.Numbers of misassemblies of different kinds (inversions, relocations, translocations, interspecies translocations (metaQUAST only) or local).Number of predicted genes, discovered either by GeneMark.hmm (for prokaryotes), GeneMark-ES or GlimmerHMM (for eukaryotes),.N50 (length of a contig, such that all the contigs of at least the same length together cover at least 50% of the assembly).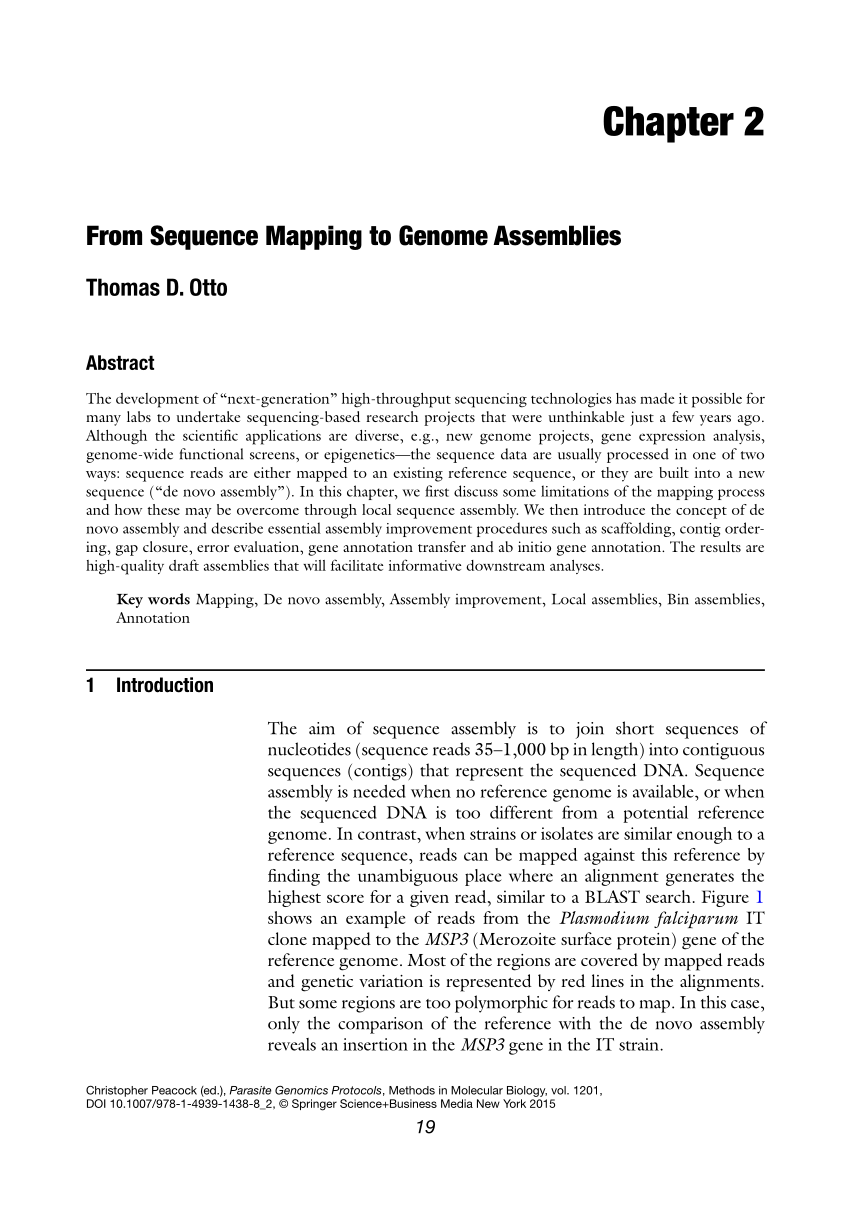 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
